Large language models are impressive at generating text, but they have a fundamental limitation: they only know what was in their training data. Ask an LLM about your internal documentation, your company's runbook procedures, or your infrastructure's specific configuration, and it will either admit ignorance or — worse — hallucinate a plausible-sounding answer that is completely wrong. Retrieval-Augmented Generation (RAG) solves this by giving the LLM access to your actual documents at query time. Instead of relying on memorized training data, the model retrieves relevant passages from your documentation and uses them as context for generating its response.
A self-hosted RAG pipeline keeps everything on your network. Your documentation never leaves your infrastructure. The embeddings (vector representations of your documents) are stored in a local database. The LLM runs on your own GPU. The result is an AI assistant that can answer questions about your specific environment — your Ansible playbooks, your architecture decisions, your incident postmortems, your internal wikis — with citations pointing to the source documents.
This guide builds the complete pipeline from scratch: document ingestion from multiple sources (Markdown, PDF, HTML, plain text), text chunking strategies that preserve context, embedding generation using a local model, vector storage with ChromaDB, retrieval-augmented query processing with Ollama, and a simple web interface for interactive use. Every component runs locally on Linux. For the foundational setup, see our complete Ollama installation guide. For GPU driver setup, see our NVIDIA driver and CUDA installation guide.
Architecture and Data Flow
# RAG Pipeline Architecture
#
# Document Ingestion:
# Files (MD, PDF, TXT, HTML) --> Chunker --> Embedding Model --> ChromaDB
#
# Query Processing:
# User Question --> Embedding Model --> ChromaDB (similarity search)
# |
# Top-K relevant chunks
# |
# Prompt Assembly:
# "Context: [chunks]
# Question: [user question]"
# |
# Ollama (LLM)
# |
# Answer with citationsInstalling the Components
Ollama (LLM and Embedding Model)
# Install Ollama
curl -fsSL https://ollama.com/install.sh | sh
sudo systemctl enable --now ollama
# Pull a capable chat model
ollama pull llama3.1:8b
# Pull a dedicated embedding model
# nomic-embed-text is small (274M params) and produces quality embeddings
ollama pull nomic-embed-text
# Verify both models are available
ollama listChromaDB (Vector Database)
# Create a virtual environment for the RAG pipeline
python3 -m venv /opt/rag-pipeline
source /opt/rag-pipeline/bin/activate
# Install ChromaDB and other dependencies
pip install chromadb sentence-transformers requests
# Install document processing libraries
pip install pymupdf python-docx beautifulsoup4 markdown
pip install tiktoken # For accurate token counting
# Verify ChromaDB works
python3 -c "import chromadb; client = chromadb.PersistentClient(path='/tmp/test_chroma'); print('ChromaDB OK')"Document Ingestion Pipeline
The ingestion pipeline reads documents from various formats, splits them into chunks, generates embeddings, and stores everything in ChromaDB. The chunking strategy is critical — chunks that are too large dilute the relevance signal, while chunks that are too small lose context.
Document Loader
#!/usr/bin/env python3
"""document_loader.py — Load documents from various formats."""
from pathlib import Path
from typing import List, Dict
import json
def load_markdown(file_path: Path) -> str:
"""Load a Markdown file and return raw text."""
return file_path.read_text(encoding="utf-8")
def load_pdf(file_path: Path) -> str:
"""Extract text from a PDF file using PyMuPDF."""
import fitz # pymupdf
doc = fitz.open(str(file_path))
text = ""
for page in doc:
text += page.get_text() + "\n"
doc.close()
return text
def load_html(file_path: Path) -> str:
"""Extract text from an HTML file."""
from bs4 import BeautifulSoup
html = file_path.read_text(encoding="utf-8")
soup = BeautifulSoup(html, "html.parser")
# Remove script and style elements
for element in soup(["script", "style", "nav", "footer"]):
element.decompose()
return soup.get_text(separator="\n", strip=True)
def load_text(file_path: Path) -> str:
"""Load a plain text file."""
return file_path.read_text(encoding="utf-8")
LOADERS = {
".md": load_markdown,
".markdown": load_markdown,
".pdf": load_pdf,
".html": load_html,
".htm": load_html,
".txt": load_text,
".text": load_text,
".rst": load_text,
".yaml": load_text,
".yml": load_text,
".conf": load_text,
".cfg": load_text,
".json": load_text,
}
def load_document(file_path: Path) -> Dict:
"""Load a document and return its content with metadata."""
suffix = file_path.suffix.lower()
loader = LOADERS.get(suffix)
if loader is None:
raise ValueError(f"Unsupported file type: {suffix}")
content = loader(file_path)
return {
"content": content,
"metadata": {
"source": str(file_path),
"filename": file_path.name,
"filetype": suffix,
"size_bytes": file_path.stat().st_size,
}
}
def load_directory(dir_path: Path, recursive: bool = True) -> List[Dict]:
"""Load all supported documents from a directory."""
documents = []
pattern = "**/*" if recursive else "*"
for file_path in sorted(dir_path.glob(pattern)):
if file_path.is_file() and file_path.suffix.lower() in LOADERS:
try:
doc = load_document(file_path)
documents.append(doc)
print(f" Loaded: {file_path.name} ({len(doc['content'])} chars)")
except Exception as e:
print(f" Error loading {file_path.name}: {e}")
return documentsText Chunking
#!/usr/bin/env python3
"""chunker.py — Split documents into overlapping chunks."""
from typing import List, Dict
import re
def chunk_text(
text: str,
chunk_size: int = 512,
chunk_overlap: int = 50,
metadata: Dict = None
) -> List[Dict]:
"""Split text into overlapping chunks, respecting sentence boundaries."""
# Clean up whitespace
text = re.sub(r'\n{3,}', '\n\n', text)
text = re.sub(r' {2,}', ' ', text)
# Split into sentences (approximate)
sentences = re.split(r'(?<=[.!?])\s+', text)
chunks = []
current_chunk = []
current_length = 0
for sentence in sentences:
sentence_length = len(sentence.split())
if current_length + sentence_length > chunk_size and current_chunk:
# Save current chunk
chunk_text_joined = " ".join(current_chunk)
chunk_metadata = dict(metadata) if metadata else {}
chunk_metadata["chunk_index"] = len(chunks)
chunk_metadata["chunk_size"] = current_length
chunks.append({
"text": chunk_text_joined,
"metadata": chunk_metadata,
})
# Keep overlap sentences for the next chunk
overlap_words = 0
overlap_start = len(current_chunk)
for i in range(len(current_chunk) - 1, -1, -1):
overlap_words += len(current_chunk[i].split())
if overlap_words >= chunk_overlap:
overlap_start = i
break
current_chunk = current_chunk[overlap_start:]
current_length = sum(len(s.split()) for s in current_chunk)
current_chunk.append(sentence)
current_length += sentence_length
# Save the last chunk
if current_chunk:
chunk_text_joined = " ".join(current_chunk)
chunk_metadata = dict(metadata) if metadata else {}
chunk_metadata["chunk_index"] = len(chunks)
chunk_metadata["chunk_size"] = current_length
chunks.append({
"text": chunk_text_joined,
"metadata": chunk_metadata,
})
return chunksEmbedding Generation and Storage
Embeddings are vector representations of text that capture semantic meaning. Similar texts produce similar vectors, which is how the retrieval step finds relevant passages. Ollama can generate embeddings using dedicated embedding models.
Embedding with Ollama
#!/usr/bin/env python3
"""embedder.py — Generate embeddings using Ollama."""
import requests
from typing import List
OLLAMA_API = "http://localhost:11434/api/embeddings"
EMBED_MODEL = "nomic-embed-text"
def get_embedding(text: str, model: str = EMBED_MODEL) -> List[float]:
"""Generate an embedding vector for a text using Ollama."""
response = requests.post(OLLAMA_API, json={
"model": model,
"prompt": text,
}, timeout=30)
if response.status_code == 200:
return response.json()["embedding"]
raise RuntimeError(f"Embedding failed: {response.status_code}")
def get_embeddings_batch(texts: List[str], model: str = EMBED_MODEL) -> List[List[float]]:
"""Generate embeddings for multiple texts."""
embeddings = []
for i, text in enumerate(texts):
if i % 50 == 0 and i > 0:
print(f" Embedded {i}/{len(texts)} chunks...")
embeddings.append(get_embedding(text, model))
return embeddingsChromaDB Vector Store
#!/usr/bin/env python3
"""vector_store.py — ChromaDB vector storage and retrieval."""
import chromadb
from typing import List, Dict, Optional
class VectorStore:
def __init__(self, persist_dir: str = "/var/lib/rag/chromadb"):
self.client = chromadb.PersistentClient(path=persist_dir)
def get_or_create_collection(self, name: str = "documents"):
"""Get or create a ChromaDB collection."""
return self.client.get_or_create_collection(
name=name,
metadata={"hnsw:space": "cosine"} # Cosine similarity
)
def add_documents(self, chunks: List[Dict], embeddings: List[List[float]],
collection_name: str = "documents"):
"""Add document chunks with embeddings to the vector store."""
collection = self.get_or_create_collection(collection_name)
ids = [f"chunk_{collection.count() + i}" for i in range(len(chunks))]
documents = [chunk["text"] for chunk in chunks]
metadatas = [chunk["metadata"] for chunk in chunks]
collection.add(
ids=ids,
documents=documents,
embeddings=embeddings,
metadatas=metadatas,
)
print(f" Added {len(chunks)} chunks to collection '{collection_name}'")
print(f" Total documents in collection: {collection.count()}")
def query(self, query_embedding: List[float], n_results: int = 5,
collection_name: str = "documents",
where: Optional[Dict] = None) -> Dict:
"""Query the vector store for similar documents."""
collection = self.get_or_create_collection(collection_name)
results = collection.query(
query_embeddings=[query_embedding],
n_results=n_results,
where=where,
include=["documents", "metadatas", "distances"],
)
return results
def get_stats(self, collection_name: str = "documents") -> Dict:
"""Get collection statistics."""
collection = self.get_or_create_collection(collection_name)
return {
"collection": collection_name,
"document_count": collection.count(),
}The RAG Query Engine
The query engine ties everything together: it takes a user question, finds relevant document chunks, builds a prompt with context, and sends it to the LLM for answer generation.
#!/usr/bin/env python3
"""rag_engine.py — Retrieval-Augmented Generation query engine."""
import requests
from typing import List, Dict, Optional
from embedder import get_embedding
from vector_store import VectorStore
OLLAMA_API = "http://localhost:11434/api/generate"
CHAT_MODEL = "llama3.1:8b"
class RAGEngine:
def __init__(self, persist_dir: str = "/var/lib/rag/chromadb",
chat_model: str = CHAT_MODEL):
self.store = VectorStore(persist_dir)
self.chat_model = chat_model
def query(self, question: str, n_context: int = 5,
collection: str = "documents") -> Dict:
"""Answer a question using RAG."""
# Step 1: Embed the question
question_embedding = get_embedding(question)
# Step 2: Retrieve relevant chunks
results = self.store.query(
query_embedding=question_embedding,
n_results=n_context,
collection_name=collection,
)
documents = results["documents"][0] if results["documents"] else []
metadatas = results["metadatas"][0] if results["metadatas"] else []
distances = results["distances"][0] if results["distances"] else []
# Step 3: Build the context-aware prompt
context_parts = []
sources = []
for i, (doc, meta, dist) in enumerate(zip(documents, metadatas, distances)):
source = meta.get("source", "unknown")
context_parts.append(f"[Source {i+1}: {meta.get('filename', 'unknown')}]\n{doc}")
sources.append({
"file": source,
"filename": meta.get("filename", "unknown"),
"chunk_index": meta.get("chunk_index", 0),
"relevance": 1 - dist, # Convert distance to similarity
})
context = "\n\n".join(context_parts)
prompt = (
"You are a helpful assistant that answers questions based on the "
"provided documentation context. Use ONLY the information from the "
"context below to answer. If the context does not contain enough "
"information to answer the question, say so clearly. "
"Always cite the source number [Source N] when using information "
"from a specific document.\n\n"
f"Context:\n{context}\n\n"
f"Question: {question}\n\n"
"Answer:"
)
# Step 4: Generate the answer
response = requests.post(OLLAMA_API, json={
"model": self.chat_model,
"prompt": prompt,
"stream": False,
"options": {
"temperature": 0.3,
"num_predict": 2048,
}
}, timeout=120)
answer = response.json().get("response", "Failed to generate response.")
return {
"question": question,
"answer": answer,
"sources": sources,
"context_chunks": len(documents),
}Complete Pipeline: Ingest and Query
#!/usr/bin/env python3
"""rag_cli.py — Command-line interface for the RAG pipeline."""
import argparse
import json
import sys
from pathlib import Path
from document_loader import load_directory, load_document
from chunker import chunk_text
from embedder import get_embeddings_batch
from vector_store import VectorStore
from rag_engine import RAGEngine
def cmd_ingest(args):
"""Ingest documents into the vector store."""
source = Path(args.source)
store = VectorStore(args.db_path)
if source.is_dir():
print(f"Loading documents from {source}...")
documents = load_directory(source, recursive=args.recursive)
elif source.is_file():
print(f"Loading {source}...")
documents = [load_document(source)]
else:
print(f"Error: {source} not found", file=sys.stderr)
sys.exit(1)
print(f"Loaded {len(documents)} documents")
# Chunk all documents
all_chunks = []
for doc in documents:
chunks = chunk_text(
doc["content"],
chunk_size=args.chunk_size,
chunk_overlap=args.overlap,
metadata=doc["metadata"],
)
all_chunks.extend(chunks)
print(f"Created {len(all_chunks)} chunks")
# Generate embeddings
print("Generating embeddings...")
texts = [c["text"] for c in all_chunks]
embeddings = get_embeddings_batch(texts)
# Store in ChromaDB
store.add_documents(all_chunks, embeddings, args.collection)
stats = store.get_stats(args.collection)
print(f"Done. Collection now has {stats['document_count']} chunks.")
def cmd_query(args):
"""Query the RAG pipeline."""
engine = RAGEngine(args.db_path, args.model)
question = " ".join(args.question)
result = engine.query(
question=question,
n_context=args.context_chunks,
collection=args.collection,
)
print(f"\nQuestion: {result['question']}\n")
print(f"Answer:\n{result['answer']}\n")
print(f"Sources ({len(result['sources'])}):")
for s in result["sources"]:
print(f" - {s['filename']} (chunk {s['chunk_index']}, "
f"relevance: {s['relevance']:.2%})")
def cmd_stats(args):
"""Show vector store statistics."""
store = VectorStore(args.db_path)
stats = store.get_stats(args.collection)
print(json.dumps(stats, indent=2))
if __name__ == "__main__":
parser = argparse.ArgumentParser(description="Self-hosted RAG Pipeline")
parser.add_argument("--db-path", default="/var/lib/rag/chromadb")
parser.add_argument("--collection", default="documents")
sub = parser.add_subparsers(dest="command", required=True)
ing = sub.add_parser("ingest", help="Ingest documents")
ing.add_argument("source", help="File or directory to ingest")
ing.add_argument("--chunk-size", type=int, default=512)
ing.add_argument("--overlap", type=int, default=50)
ing.add_argument("--recursive", action="store_true", default=True)
qry = sub.add_parser("query", help="Query the knowledge base")
qry.add_argument("question", nargs="+")
qry.add_argument("-n", "--context-chunks", type=int, default=5)
qry.add_argument("-m", "--model", default="llama3.1:8b")
st = sub.add_parser("stats", help="Show collection stats")
args = parser.parse_args()
{"ingest": cmd_ingest, "query": cmd_query, "stats": cmd_stats}[args.command](args)Using the Pipeline
# Create the data directory
sudo mkdir -p /var/lib/rag/chromadb
sudo chown $USER:$USER /var/lib/rag -R
# Ingest your documentation
python3 rag_cli.py ingest /path/to/your/documentation/ --recursive
# Ingest a specific file
python3 rag_cli.py ingest /path/to/runbook.md
# Query your knowledge base
python3 rag_cli.py query "How do I restart the production database cluster?"
python3 rag_cli.py query "What is our backup retention policy?"
python3 rag_cli.py query "What monitoring alerts exist for the payment service?"
# Check collection statistics
python3 rag_cli.py statsImproving Retrieval Quality
The quality of RAG output depends primarily on retrieval quality — finding the right document chunks for each question. Several techniques improve retrieval beyond basic similarity search.
Hybrid Search: Combine Vector and Keyword Search
# ChromaDB supports filtering by metadata, but for true hybrid search,
# combine vector similarity with BM25 keyword matching.
pip install rank-bm25
from rank_bm25 import BM25Okapi
def hybrid_search(question, store, n_results=5):
"""Combine vector similarity with BM25 keyword search."""
# Vector search
question_embedding = get_embedding(question)
vector_results = store.query(question_embedding, n_results=n_results * 2)
# BM25 search on the same collection
collection = store.get_or_create_collection()
all_docs = collection.get(include=["documents"])
tokenized_docs = [doc.lower().split() for doc in all_docs["documents"]]
bm25 = BM25Okapi(tokenized_docs)
bm25_scores = bm25.get_scores(question.lower().split())
# Combine scores (normalize and weight)
# 0.7 weight for vector similarity, 0.3 for BM25
combined = {}
for i, (doc_id, dist) in enumerate(
zip(vector_results["ids"][0], vector_results["distances"][0])
):
combined[doc_id] = 0.7 * (1 - dist)
for i, score in enumerate(bm25_scores):
doc_id = all_docs["ids"][i]
if doc_id in combined:
combined[doc_id] += 0.3 * (score / max(bm25_scores))
else:
combined[doc_id] = 0.3 * (score / max(bm25_scores))
# Return top N results
sorted_results = sorted(combined.items(), key=lambda x: x[1], reverse=True)
return sorted_results[:n_results]Query Rewriting for Better Retrieval
# Use the LLM to rewrite ambiguous queries before searching
def rewrite_query(original_query, model="llama3.1:8b"):
"""Use the LLM to expand or clarify a search query."""
prompt = (
"Rewrite this search query to be more specific and include "
"relevant technical terms. Output only the rewritten query, "
"nothing else.\n\n"
f"Original query: {original_query}\n"
"Rewritten query:"
)
response = requests.post(OLLAMA_API, json={
"model": model,
"prompt": prompt,
"stream": False,
"options": {"temperature": 0.1, "num_predict": 100}
})
rewritten = response.json().get("response", original_query).strip()
return rewritten
# Example:
# "how to fix the database" might be rewritten to:
# "database troubleshooting procedures connection errors performance issues PostgreSQL MySQL"Running as a Service
# systemd service for the RAG API
sudo tee /etc/systemd/system/rag-pipeline.service <<'EOF'
[Unit]
Description=RAG Pipeline API Service
After=network.target ollama.service
[Service]
Type=simple
User=rag
Group=rag
WorkingDirectory=/opt/rag-pipeline
ExecStart=/opt/rag-pipeline/bin/python3 rag_api.py
Restart=on-failure
RestartSec=5
Environment=RAG_DB_PATH=/var/lib/rag/chromadb
Environment=OLLAMA_API=http://localhost:11434
NoNewPrivileges=true
ProtectSystem=strict
ProtectHome=true
ReadWritePaths=/var/lib/rag
PrivateTmp=true
[Install]
WantedBy=multi-user.target
EOF
sudo useradd -r -s /usr/sbin/nologin rag
sudo chown -R rag:rag /var/lib/rag
sudo systemctl daemon-reload
sudo systemctl enable --now rag-pipeline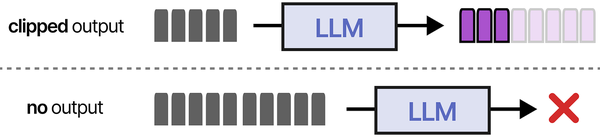
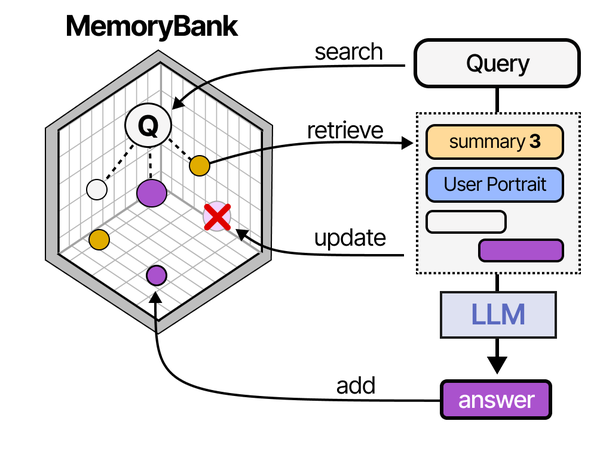
Building a self-hosted RAG pipeline on Linux with Ollama implements the retrieval-augmented generation architecture that Grootendorst and Alammar detail extensively in An Illustrated Guide to AI Agents. Their RAG pipeline diagram shows the complete flow: documents are chunked and embedded into a vector store, user queries are similarly embedded, relevant chunks are retrieved via similarity search, and the retrieved context is prepended to the prompt before generation. The MemoryBank architecture they describe extends this with persistent knowledge management across sessions. Running this entire pipeline locally with Ollama for inference and ChromaDB or Qdrant for vector storage gives Linux administrators full control over their organization's knowledge retrieval system.
Related Articles
- AnythingLLM on Linux: Self-Hosted RAG with Ollama and Document Chat
- AI Document OCR on Linux: Open Source Pipeline with Tesseract and LLMs
- Ollama and LangChain on Linux: Build AI Agents with Local Models
- Self-Hosted AI Translation Server on Linux with Ollama
Further Reading
- An Illustrated Guide to AI Agents by Maarten Grootendorst and Jay Alammar — Visual guide to agent memory, tools, and reasoning.
- LLMs in Production by Christopher Brousseau and Matthew Sharp — Practical deployment of language models from training to production.
- Agentic AI in Enterprise by Sumit Ranjan, Divya Chembachere, and Lanwin Lobo — Enterprise architecture patterns for agentic AI systems.
Frequently Asked Questions
How much documentation can the pipeline handle?
ChromaDB on a single server handles up to about 1 million vector chunks comfortably. With a chunk size of 512 tokens, that represents roughly 500,000 pages of documentation — far more than most organizations have. Ingestion speed depends on the embedding model: nomic-embed-text through Ollama processes about 50-100 chunks per second on a GPU. A 10,000-page documentation set takes about 15-30 minutes to ingest.
Which embedding model should I use?
For English documentation, nomic-embed-text through Ollama offers an excellent balance of quality and speed. It produces 768-dimensional vectors and runs quickly even on modest hardware. For multilingual documentation, mxbai-embed-large or snowflake-arctic-embed handle multiple languages well. The embedding model has a larger impact on retrieval quality than the chat model has on answer quality — invest in the best embedding model your hardware can support.
How do I update documents after they change?
The simplest approach is to re-ingest changed documents. Delete the old chunks for the specific file using ChromaDB's metadata filtering (collection.delete(where={"source": "/path/to/file.md"})), then re-ingest the updated file. For large documentation sets that change frequently, implement incremental ingestion that tracks file modification times and only re-processes changed files.
Why does the RAG pipeline sometimes return irrelevant context?
Three common causes: chunk size is too large (dilutes relevance), the embedding model does not capture domain-specific terminology well, or the query is too vague. Start by reducing chunk size (try 256 tokens instead of 512). If that does not help, try query rewriting to expand abbreviated or ambiguous questions. Finally, consider whether your documents need preprocessing — tables, code blocks, and bullet lists sometimes embed poorly as raw text.
Can I use this with Open WebUI instead of the command line?
Yes. Open WebUI has built-in RAG support. You can upload documents through its web interface, and it handles chunking, embedding, and retrieval internally. It uses the Ollama embedding endpoint and stores vectors in its own database. The pipeline described in this article gives you more control over chunking strategies, embedding models, and retrieval algorithms, but Open WebUI provides a simpler path if you want RAG without writing code.


